Qiuyu Fu made important contribution to this study.
Hong Lin, Xiaozhe Zhang, Li Liu, Qiuyu Fu, Chuanlong Zang, Yan Ding, Yang Su, Zhixue Xu, Sining He, Xiaoli Yang, Xiayun Wei, Haibin Mao, Yasong Cui, Yi Wei, Chuanzheng Zhou, Lilin Du, Niu Huang, Ning Zheng, Tao Wang, and Feng Rao
Abstract
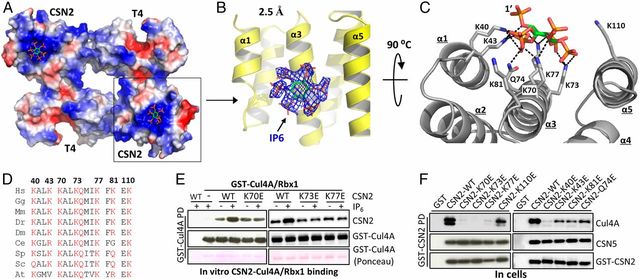
The Cullin-RING ligases (CRLs) are the largest family of ubiquitin E3s activated by neddylation and regulated by the deneddylase COP9 signalosome (CSN). The inositol polyphosphate metabolites promote the formation of CRL–CSN complexes, but with unclear mechanism of action. Here, we provide structural and genetic evidence supporting inositol hexakisphosphate (IP6) as a general CSN cofactor recruiting CRLs. We determined the crystal structure of IP6 in complex with CSN subunit 2 (CSN2), based on which we identified the IP6-corresponding electron density in the cryoelectron microscopy map of a CRL4A–CSN complex. IP6 binds to a cognate pocket formed by conserved lysine residues from CSN2 and Rbx1/Roc1, thereby strengthening CRL–CSN interactions to dislodge the E2 CDC34/UBE2R from CRL and to promote CRL deneddylation. IP6 binding-deficient Csn2K70E/K70E knockin mice are embryonic lethal. The same mutation disabled Schizosaccharomyces pombe Csn2 from rescuing UV-hypersensitivity of csn2-null yeast. These data suggest that CRL transition from the E2-bound active state to the CSN-bound sequestered state is critically assisted by an interfacial IP6 small molecule, whose metabolism may be coupled to CRL–CSN complex dynamics.
PNAS first published February 11, 2020 https://doi.org/10.1073/pnas.1911998117
